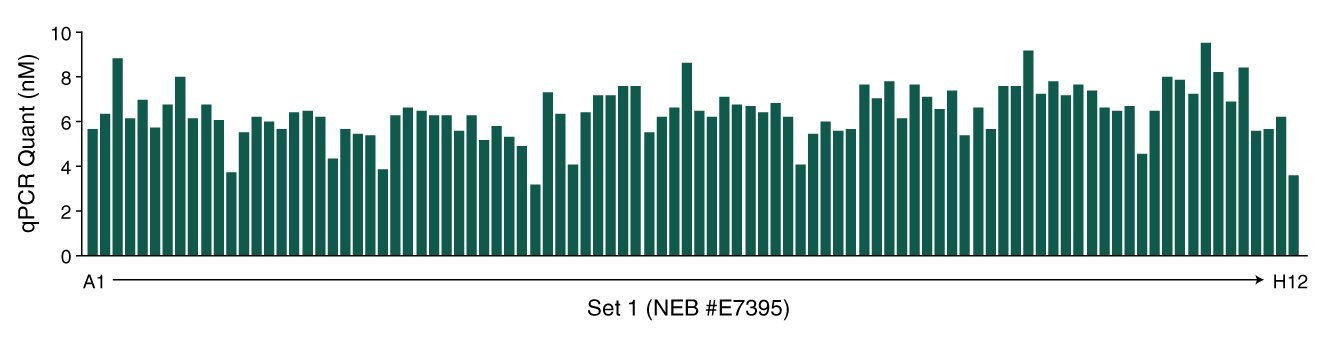CapZyme-Seq: A 5′-RNA-Seq Method for Differential Detection and Quantitation of NAD-Capped and Uncapped 5′-Triphosphate RNA - ScienceDirect

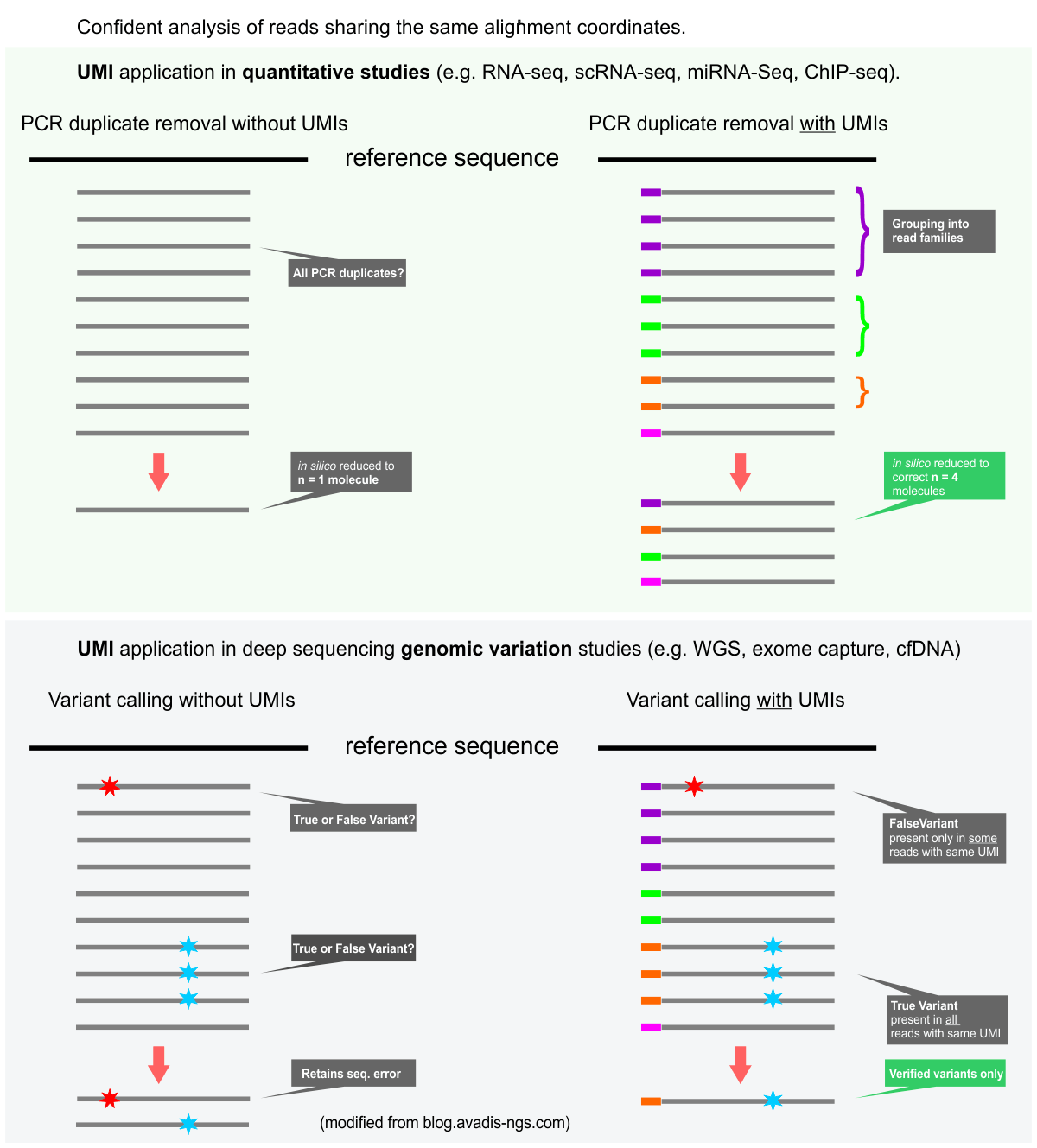
UMI incorporation into RNA-seq. a Overall workflow. Schematic of a read... | Download Scientific Diagram
![Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ] Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7755/1/fig-1-full.png)
Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]

NEBNext Multiplex Oligos for Illumina (Unique Dual Index UMI Adaptors DNA Set 1) (E7395) - New England Biolabs Japan

STARR‐seq and UMI‐STARR‐seq: Assessing Enhancer Activities for Genome‐Wide‐, High‐, and Low‐Complexity Candidate Libraries - Neumayr - 2019 - Current Protocols in Molecular Biology - Wiley Online Library

Asymmetrical barcode adapter-assisted recovery of duplicate reads and error correction strategy to detect rare mutations in circulating tumor DNA | Scientific Reports
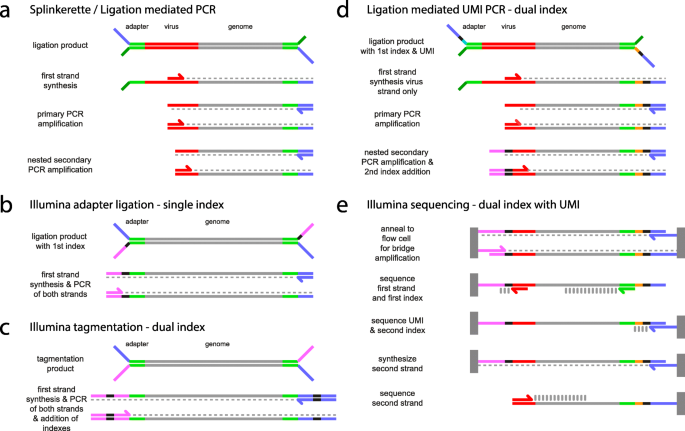
LUMI-PCR: an Illumina platform ligation-mediated PCR protocol for integration site cloning, provides molecular quantitation of integration sites | Mobile DNA | Full Text
![Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ] Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7755/1/fig-4-2x.jpg)
Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]

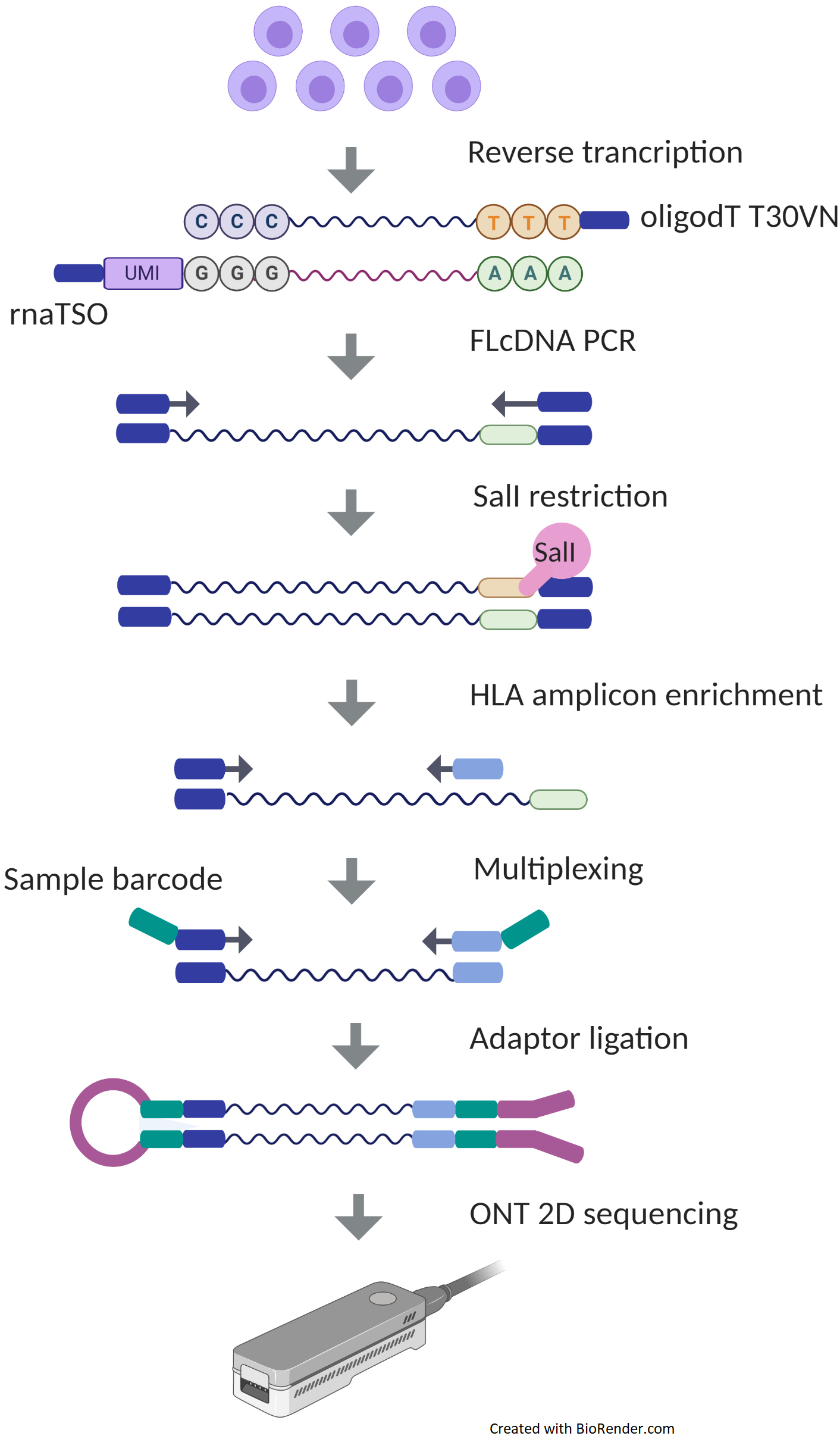
Overview of the different methods used for adapter addition to antibody... | Download Scientific Diagram

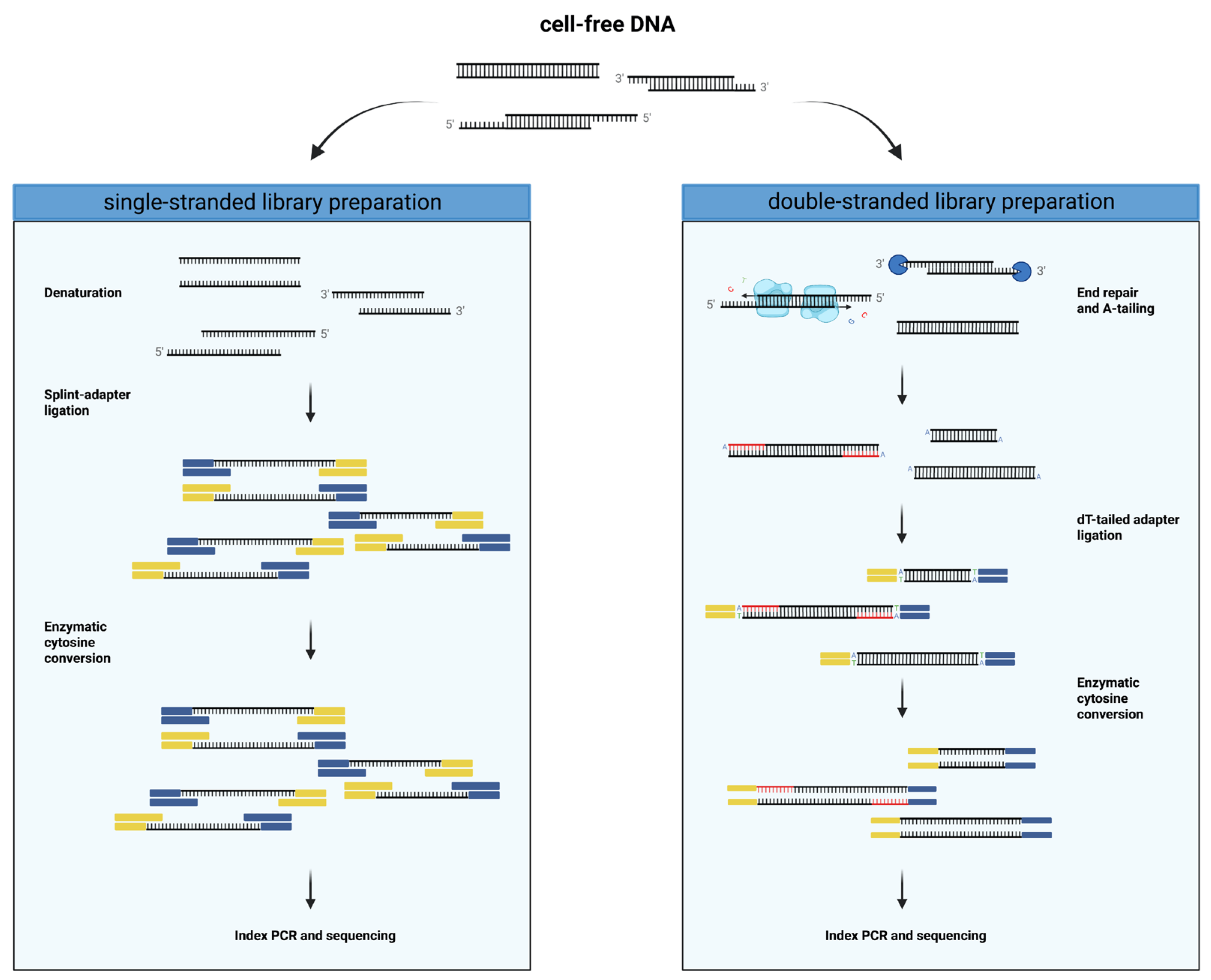
Plasma contains ultrashort single-stranded DNA in addition to nucleosomal cell-free DNA - ScienceDirect