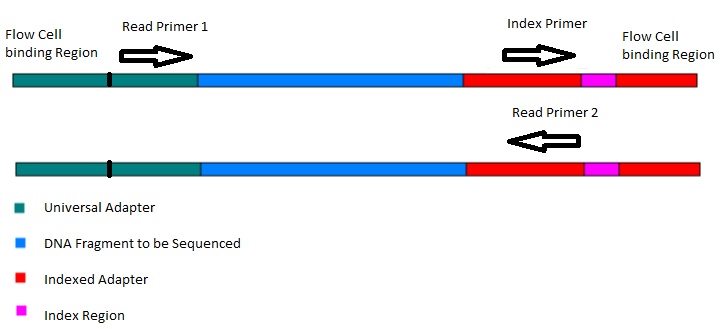
Preparation of Amplicon Libraries for Metabarcoding of Marine Eukaryotes Using Illumina MiSeq: The Adapter Ligation Method | SpringerLink

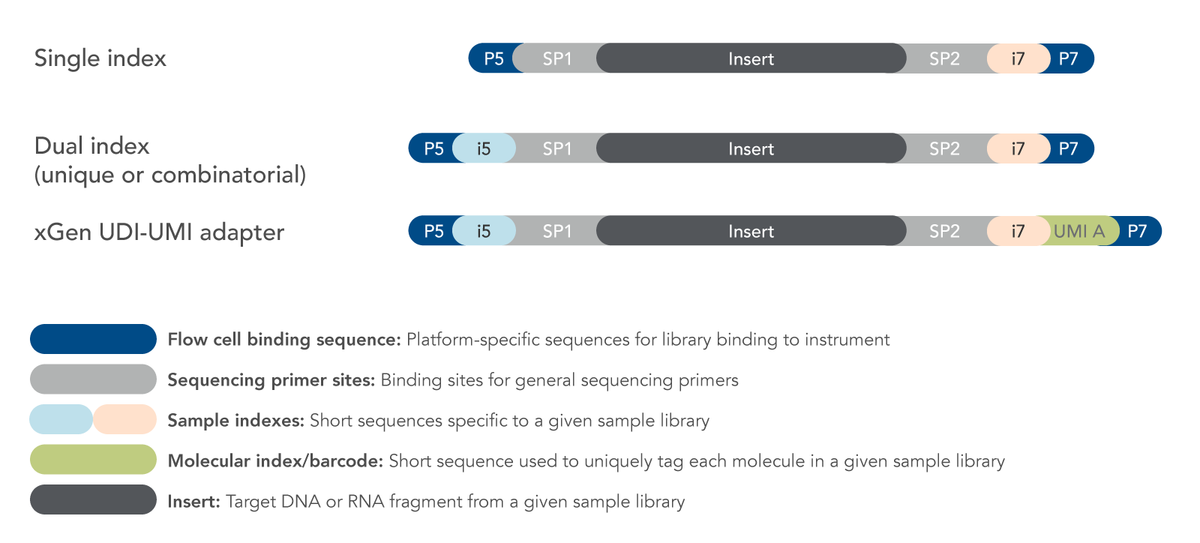
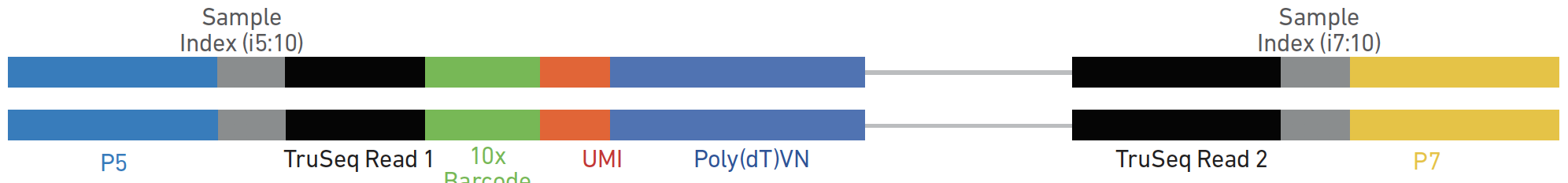
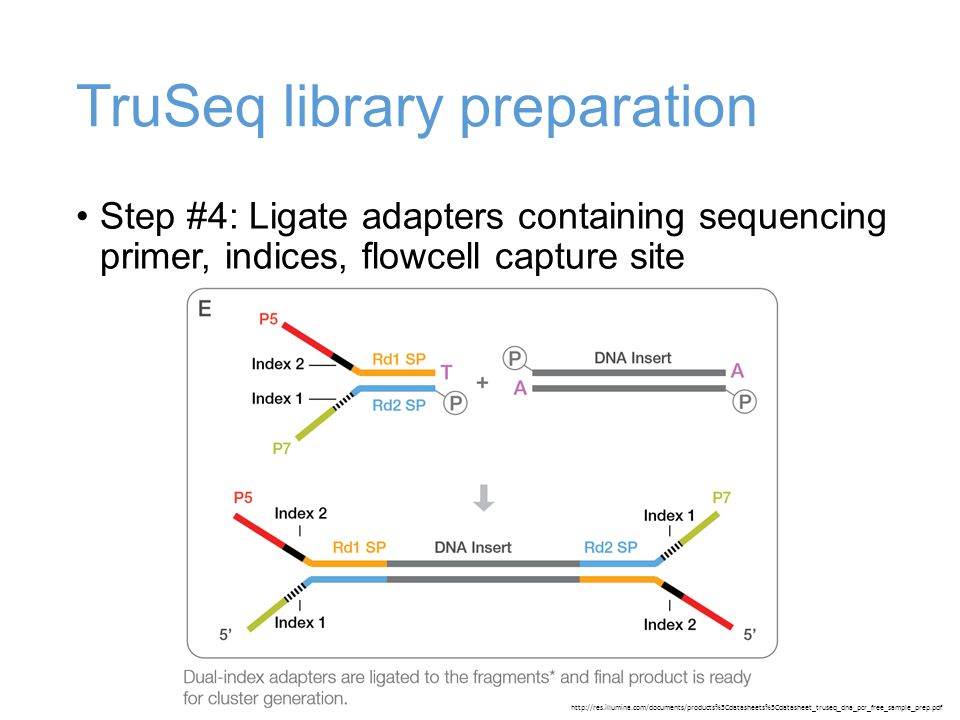
Incorporation of unique molecular identifiers in TruSeq adapters improves the accuracy of quantitative sequencing | bioRxiv

Incorporation of unique molecular identifiers (UMIs) in TruSeq adapters improves the accuracy of quantitative sequencing | RNA-Seq Blog

Incorporation of unique molecular identifiers (UMIs) in TruSeq adapters improves the accuracy of quantitative sequencing | RNA-Seq Blog
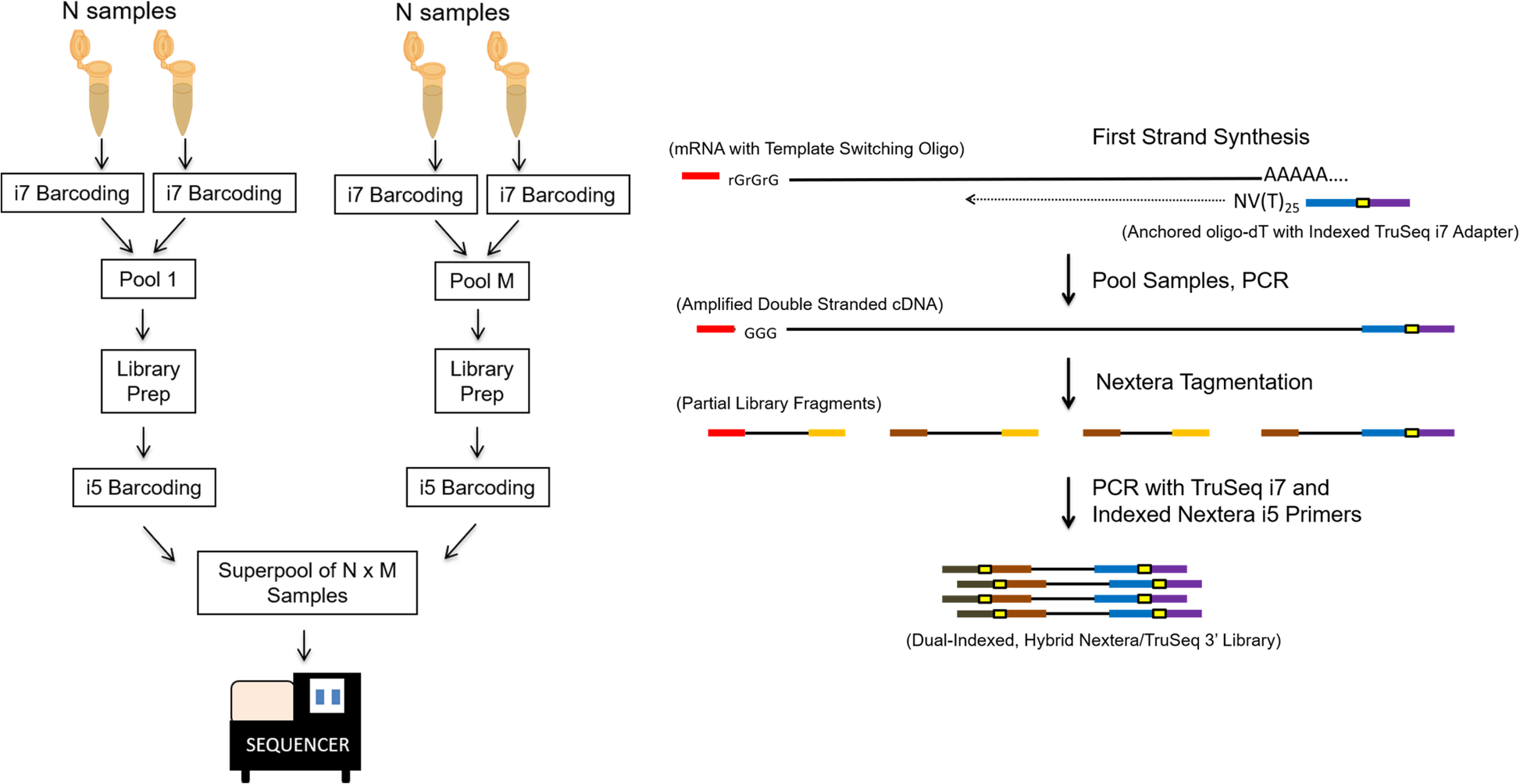
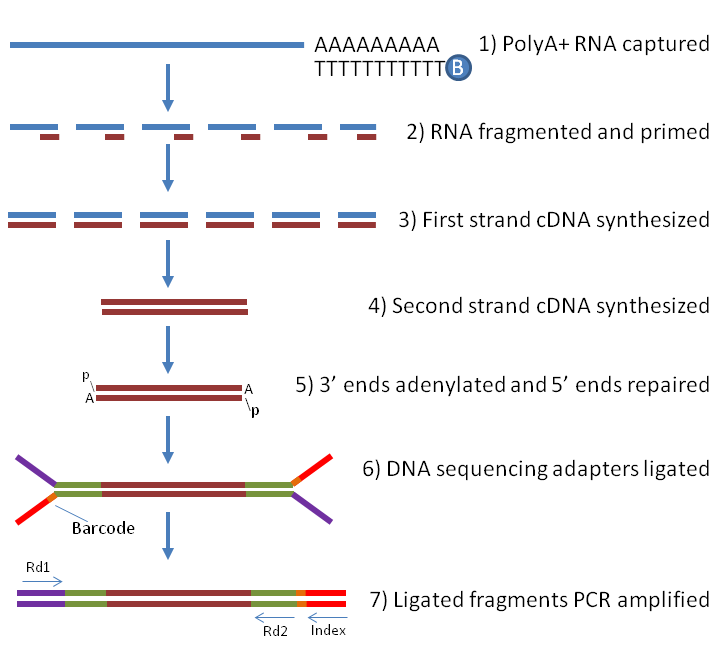
3'Pool-seq: an optimized cost-efficient and scalable method of whole-transcriptome gene expression profiling | BMC Genomics | Full Text

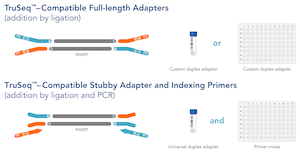
Overview of the different methods used for adapter addition to antibody... | Download Scientific Diagram
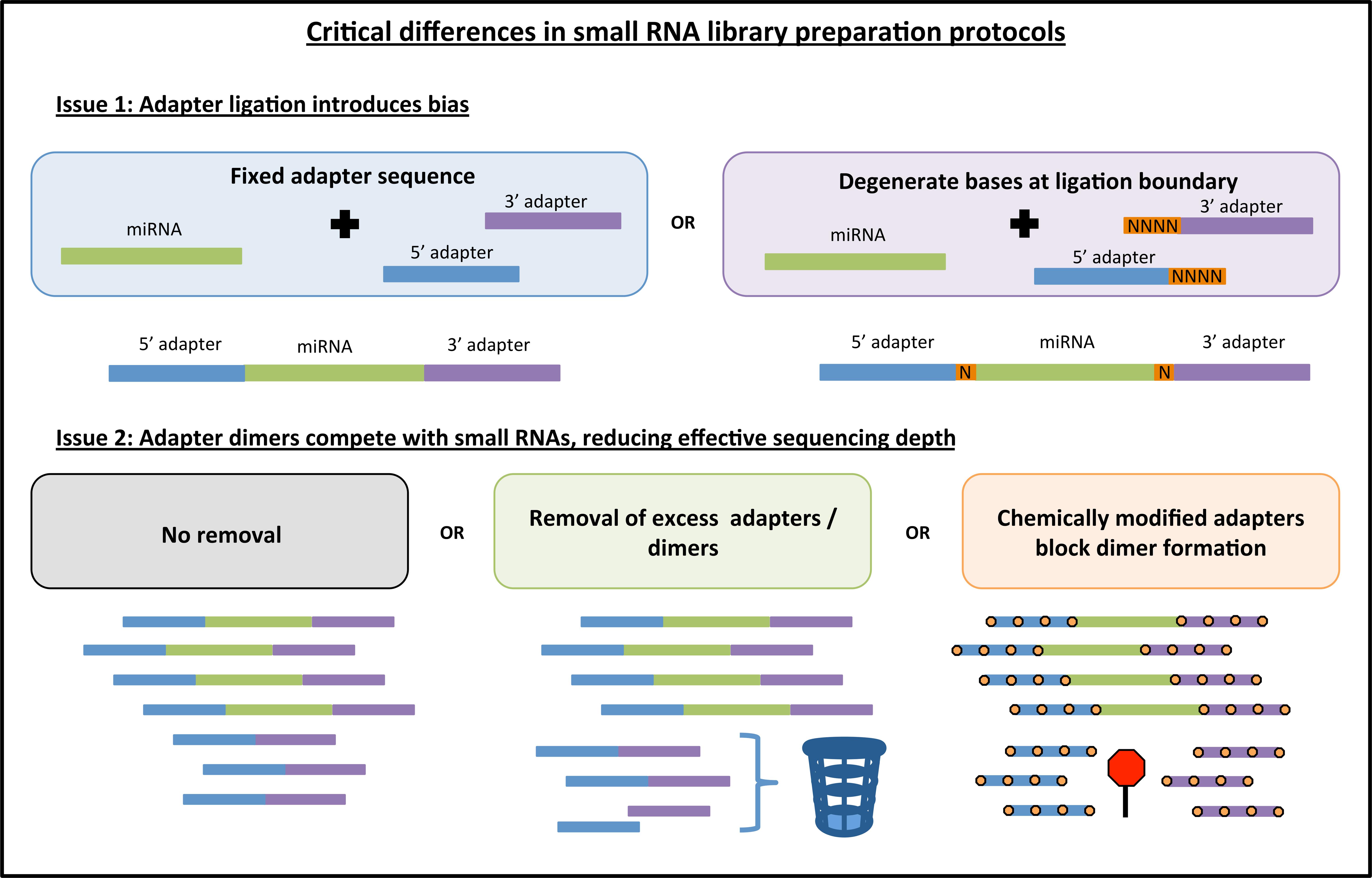
Quantitative Bias in Illumina TruSeq and a Novel Post Amplification Barcoding Strategy for Multiplexed DNA and Small RNA Deep Sequencing | PLOS ONE